Uncovering the source of CO2 that’s respired when dry soil is rewet
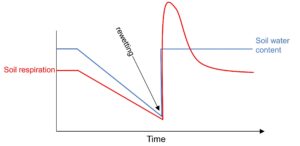
Soil respiration is one of the largest components of the global C cycle, but we do not have a good mechanistic understanding of the tens of gigatonnes per annum of respiration associated with drying followed by rewetting. In all soils drying reduces microbial activity (including respiration), while rewetting precipitates an explosive increase in microbial activity and respiration. We are actively researching the mechanistic basis for response to drying and rewetting, with a particular focus on what is fuelling the explosive pulse of CO2 when dry soil is re-wet. To uncover the sources of C that are respired we are making extensive use of mass spectrometry based metabolomics, isotope labelling and quantification of 13CO2efflux (via tuneable diode laser).
Metabolomics of plants and soil
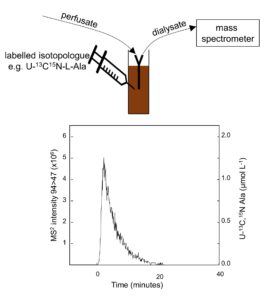
We have developed a suite of bespoke methods for analysis of plants and soil, and notched up a steady stream of success stories and firsts:
- Development of D2O-lipid-stable isotope probing to quantify the synthesis of microbial lipids in soil (Soil Biology and Biochemistry 177: 108912)
- Development of LC-MS analytical pipeline for analysis of microbial lipids in soil (Eur J Soil Sci, 69: 791-803)
- Development of D2O labelling, GC-MS and CE-MS to quantify synthesis of metabolites in soil (Soil Biology and Biochemistry 16:108543)
- Development of online microdialysis-mass spectrometry for measuring fluxes of small organic N compounds added to soil (Soil Biology and Biochemistry 123: 266-275)
- First identification and quantification of peptides in soil (Soil Biology and Biochemistry 63: 80-84)
- Development of analytical methods for comprehensive profiling of small organic N compounds in soil by CE-MS (Soil Biology and Biochemistry 57:444-450) and LC-MS (Soil Biology and Biochemistry 78: 233-242)
- Evaluation of methods for extracting organic N from the soil microbial biomass(Soil Biology and Biochemistry 81: 67-76)
- First comprehensive profiling of polar metabolites in the model tree species Eucalyptus (Metabolomics 8:186–200)
- Discovery of two metabolites responsible for osmotic adjustment in Acacia spp. that were overlooked by the previous 30+ years of research (Plant, Cell and Environment 34: 1609–1629)
- First GC-MS metabolomics mass spectral database for methane and ammonia chemical ionization (Metabolomics 9: 110-120)
Phosphorus economics of plants and soil
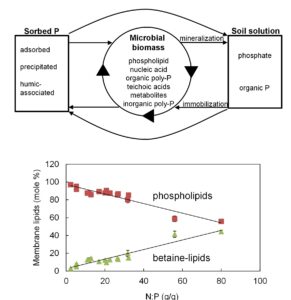
In the microbial biomass of P-poor soils (as indicated here by high N:P) P-free betaine lipids substitute for phospholipids. Data are for 3 replicate soil samples from each of 16 sites, error bars are 1SE.
Phosphorus (P) availability limits productivity across broad swathes of Earth, and is instrumental in shaping Australia’s unique flora. In natural systems P availability is largely a function of the rate that organic forms of P are converted into inorganic forms, but we know little about how microbial P compounds are mineralized. At the same time, we know plants have a raft of adaptations to P availability, but in comparison have little idea of the traits that permit soil microbial communities to flourish across broad ranges in P availability. This project is addressing three key knowledge gaps:
- To what extent do microbial communities adapt to P availability by fine-tuning cellular composition, and do microbes use similar P-saving traits as plants?
- How are microbial P compounds (e.g. nucleic acids, phospholipids, sugar phosphates, adenylates) mineralized?
- Are there feedbacks between cellular composition of microbes and phosphorus availability?
First results indicate one of the ways soil microbial communities from P-poor soils minimize cellular P requirements is by substituting phospholipids with P-free betaine lipids (see fig and Soil Biology and Biochemistry)
Ecosystem N cycling and N uptake by plants
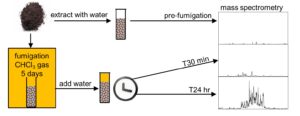
A central assumption of ecosystem N cycling has been that organic nitrogen (N) must be converted to inorganic N to be available for plant uptake, and thus N mineralization has been viewed as the bottleneck in plant N nutrition. We are investigating the hypothesis that plants can bypass the mineralization bottleneck by taking up organic N. Ongoing projects are making major contributions to our understanding of 1) limiting steps in the N cycle, 2) forms of N in soil, and 3) uptake of diverse forms of N by plants. Most recently we developed a new analytical approach to investigate the breakdown of organic N. We determined peptides were the main products of DON breakdown, and breakdown of nucleic acids and lipids also supplied significant N (see Fig and Soil Biology and Biochemistry).
